Sculpt for SGI, Windows 95/NT and PowerMac
What is it?
This review describes the new version 2.1 of Sculpt for Windows95/NT and Powermac and also covers version 1.5 for SGI (with some comments on the new version 2.0 currently in Beta testing).
So what is sculpt? In brief its a molecular modelling program that can import pdb /chemdraw files and display the molecule in a number of the common rendering styles (space fill, sticks and tubes). Minimisation takes place using a molecular mechanics force field that incorporates Van der Waals, electrostatics and hydrogen bonding.
But there's more to it...... before the reader skips to the next article thinking "who needs another MM programme", there is something different about Sculpt - a lot different. Sculpt allows the modeller to grab (virtually of course!) the model and manipulate it just as one would a plastic model. Further minimisations are carried out in real time so that there is visual feedback as to the consequences of the modification.
Sculpt really simulates the experience of playing with plastic models with the added advantage that one does not need to find enough balls and sticks of the right type and colour! For example, single atoms can be moved by simply selecting and dragging with the mouse. More or less instantly, the rest of the molecule responds. Visually this makes for a stunning presentation. In fact, my sculpt sessions regularly attracted an audience of colleagues passing through the office. This is quite a different experience to modifying a bond angle via a command line or input box and pressing the minimise button to slowly find a new (usually local minima).
Molecule/atom manipulation.
Since the prime function of sculpt is to investigate the interaction of atoms within a molecule or between molecules, the ability to control the movement/positioning of atoms/molecules is of paramount importance.
Sculpt offers a number of powerful ways to control the whole process. First, either the whole or parts of the structure can be frozen leaving the user (and the CPU) to concentrate on the critical interactions. This can be done via selection of residues/molecules from the command line (SGI) or a pull down menu (PC/Mac). Even further control can be exercised via the selection of alpha helical regions which then act as rigid moieties. Single atoms can be tugged (selected via the mouse and dragged) in any direction. Whole molecules can be grouped together so that this has the effect of moving the whole.
Tethering of two or more atoms pulls them together when auto minimise is engaged. All this adds up to an impressive degree of control over the process of, for example drug receptor docking. Few programmes can offer such ability to control the degree of rigidity/flexibility of a docking process.
Display/image manipulation facilities.

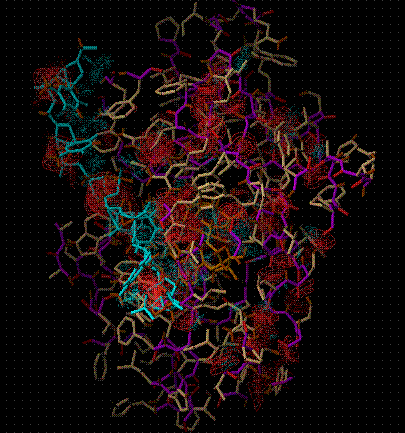
(Red wire frame represents unfavourable interaction, blue favourable)
Stick type displays are the standard sculpt display mode with tubes/cpk spheres as options. Graphics quality can be toggled to a higher resolution mode. Overall the graphics quality is adequate, but not overly impressive compared to high end modelling programmes (comparing sculpt with a range of modelling software run on the same machine).
Rotations can be carried out in all three dimension separately or combined. Translation in the Z plane is particularly useful when the perspective mode is selected combined with Z clipping. Labelling Atoms can be selected singly (resulting in the residue/atom name display), or multiply (resulting in distances/bond angle/dihedral angle display).
Drawbacks.
1. File importing
One of the most serious limitations of Sculpt is the restricted facility for importing structures from other modelling programs. Since none of the versions of Sculpt have a build facility, this is an important issue. The PC/Mac versions have the ability to paste in chemdraw structures and these then undergo a 2D to 3D conversion. All versions of Sculpt can import pdb file format. This works well if the file is genuine standard pdb format e.g. a protein crystal structure.
However problems were encountered with structures built with the hyperchem modelling program. Hyperchem numbers all atoms as if a different residue. While the atoms were then correctly imported into sculpt, no connectivity was presented. One workaround is to manually renumber the pdb file using a text editor so that each atom is the same residue. While this is not such a big job on a small molecule, renumbering a long section of DNA was a tedious task.
It would of course be unfair to blame Sculpt for this, as many modelling programs use a none standard pdb format. However, it must be borne in mind as a potential problem. Incidentally the ab initio program Gaussian 94 allows the import/output of pdb files using the newzmat conversion programme and this was successful in many cases in solving the above problem. Not many people think of Gaussian 94 as a format converter though!
2. Input of charges.
The force field employed by Sculpt offers the choice of Van der Waals and/or electrostatic interactions. The atoms charges for protein residues are taken from the standard pdb tables. Of course any other molecule needs to have charges added. These can be easily obtained from either MM or semi-empirical QM software. Inputting charges can only be done manually using a text editor, by modifying the structure saved in Sculpt's own format. As can be imagined this can be quite a task for a large molecule. Worse still, any modification to the molecule, even a change of one atom means the whole job need to be done again.
General conclusion.
A great programme that will get better. It should be in the armoury of all those interested in modelling. In addition, a role in teaching at the higher levels can be envisaged
Interactive simulations, Inc,
5330 Carrol Canyon Rd,
#203 San Diego, CA 92121,
tel 619 658 9462
fax 619 658 9463
Test machines.
Pentium 166Mhz 32Mb Ram Windows95a
PowerMac Performa 6300 8Mb Ram (+16Mb virtual) System 7.51
Silicon Graphics Indigo 2 Irix 6.2 R10000 64Mb Ram